Overview
Neurodegenerative diseases are common, debilitating, and often untreatable. We believe that a better understanding of the basic biology of these diseases will serve as a foundation upon which to develop effective therapies. Our group focuses on two related neurodegenerative diseases, frontotemporal dementia (FTD) and amyotrophic lateral sclerosis (ALS). These diseases are often inherited, and although many of the causal genes are known we don’t understand the normal function of their encoded proteins or how mutations in these genes alters upstream biological processes.
FTD and ALS

FTD is a progressive neurodegenerative disease that causes behavioral changes and language dysfunction. ALS is a neuromuscular disease that causes progressive motor decline and paralysis. Neither disease is treatable.
FTD causes loss of neurons in the frontal and temporal lobes of the cerebral cortex, while ALS causes death of motor neurons in the brain and spinal cord. Curiously, these apparently divergent diseases are related at the clinical, pathological, and genetic level. Patients with FTD frequently develop additional motor neuron degeneration, and patients with ALS often have co-existent FTD. Pathologically, FTD and ALS are often characterized by accumulations of the same aggregated proteins. Genetically, FTD and ALS can be caused by inherited mutations in the same genes. Despite remarkable progress in identifying many of the genes associated with FTD and ALS, we know relatively little about how mutations in these genes cause disease.
Identification of converging mechanisms of FTD/ALS:
Heroic efforts over the past few decades by human geneticists have identified most of the genes associated with familial forms of FTD/ALS. In parallel, technical revolutions in stem cell biology, genome engineering, and ‘omics now allow us to interrogate the function of these genes in disease-relevant cellular models. In the context of these advances, our field is poised to make giant leaps in understanding the molecular underpinnings of these related diseases.
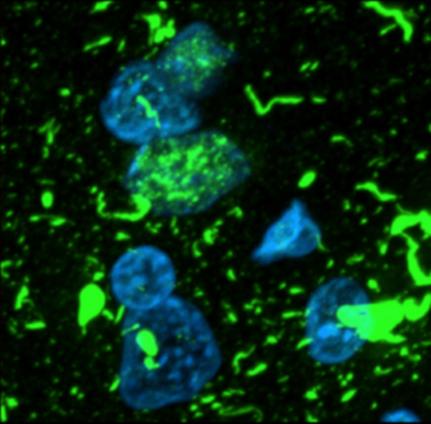
Of the roughly 40 genes associated with FTD/ALS, the majority encode proteins that appear to regulate endolysosomal biology or are components of ribonuclear protein (RNP) complexes. It is likely that at least some of these proteins regulate converging biological pathways, given the similar clinical and pathological phenotypes that ensue when mutations alter their function. For example, individuals with homozygous mutations in the GRN gene, which cause complete loss of progranulin expression, cause a lysosomal storage disease known as neuronal ceroid lipofuscinosis (NCL). We recently found that individuals with FTD-related heterozygous mutations in the GRN gene develop accumulations of pathologicAggregates of TDP-43 in the brain of an FTD patientlysosomal storage material. Progranulin localizes to the lysosome, suggesting a crucial (but undefined) role in regulation of lysosomal biology. Numerous additional FTD/ALS-related genes also regulate endolysosomal biology, including TMEM106B, which encodes a lysosomal membrane protein. Protein-coding polymorphisms in the TMEM106B gene dramatically alter the age of onset of FTD in individuals with concomitant GRN mutations. TMEM106 polymorphisms also appear to alter the clinical phenotypes of individuals with mutations in C9ORF72, another FTD/ALS-related gene that appears to regulate lysosomal function. Together, these findings strongly suggest the presence of converging molecular pathways that span multiple individual FTD/ALS genes, though the precise number, composition, and biology of such pathways remains unclear.
Research Strategy
A major focus of our lab is to systematically map how loss of function of FTD/ALS genes affects neuronal cell biology. We are using a combination of forward genetic, proteomic and transcriptomic approaches as unbiased discovery approaches to map converging and diverging pathways across multiple FTD/ALS genes in parallel. We perform these mapping studies are in human iPSC-derived neurons, using a newly-developed method that allows us to scalably generate hundreds of millions of neurons. Top pathways identified in our mapping initiatives are further investigated using traditional hypothesis-based cellular, molecular, biochemical, and imaging approaches. We anticipate that identification and pharmacological manipulation of such pathways may be particularly fruitful for development of disease modifying therapies.
